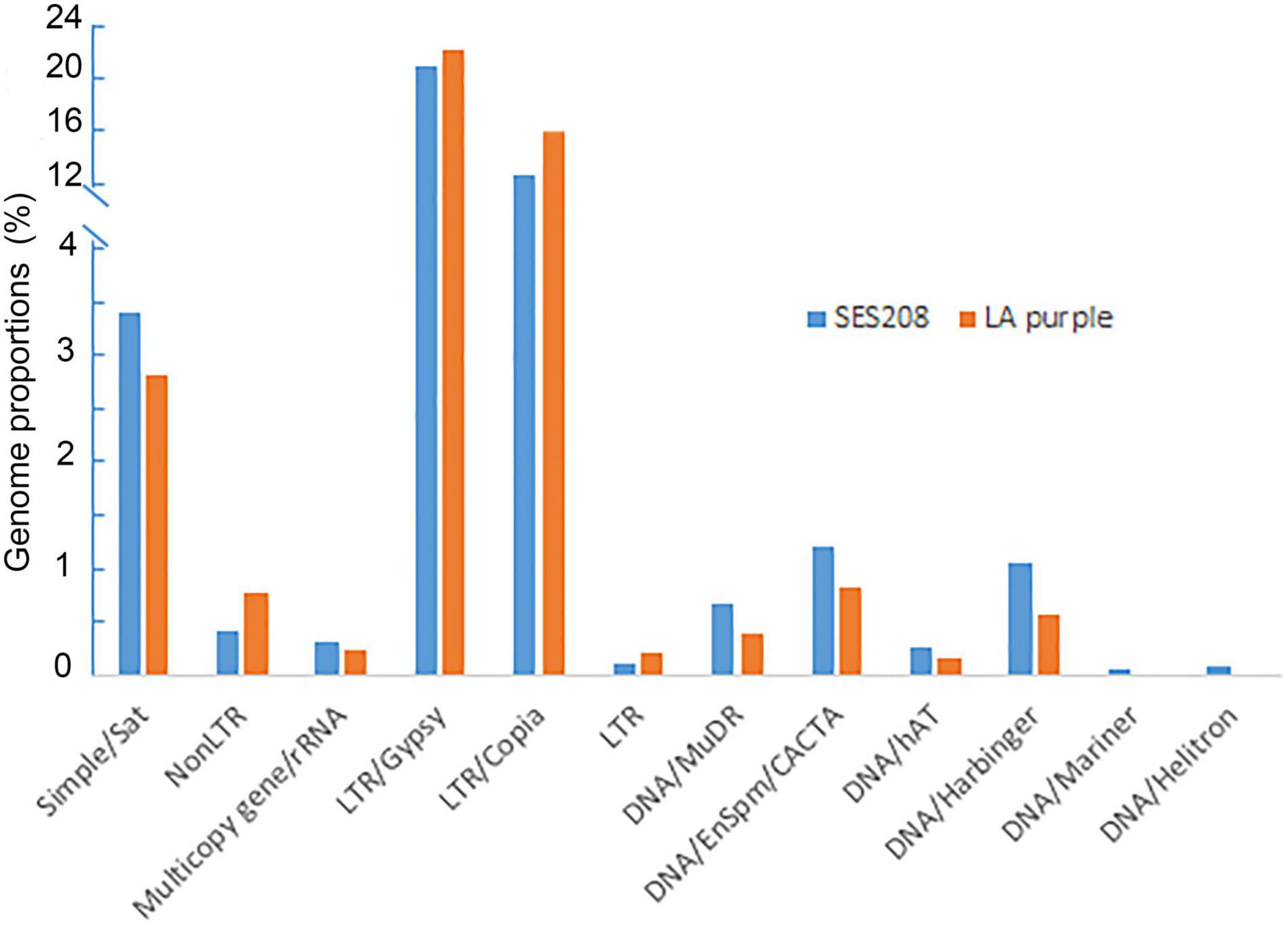
Frontiers | Characterization of Repetitive DNA in Saccharum officinarum and Saccharum spontaneum by Genome Sequencing and Cytological Assays

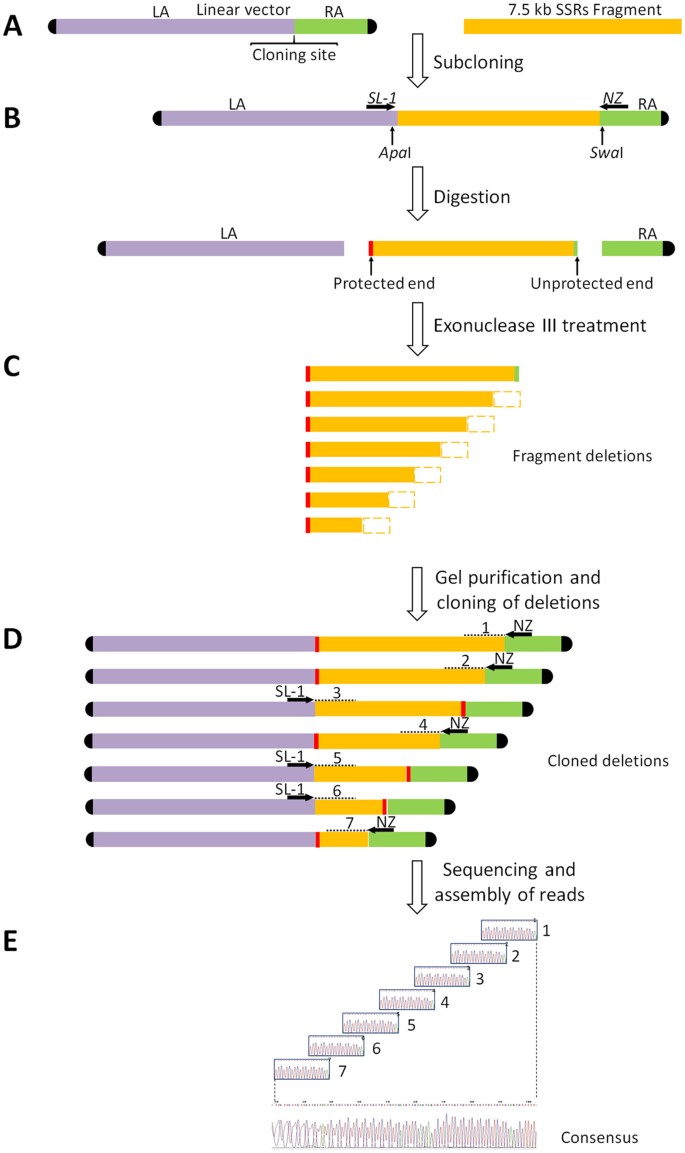
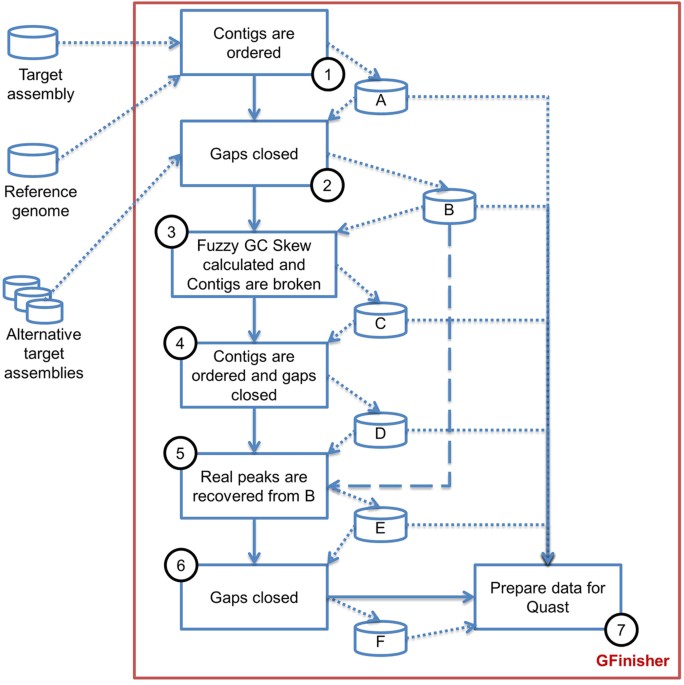
Flowchart of the sequence-assembly procedure developed in this study.... | Download Scientific Diagram

Fully phased human genome assembly without parental data using single-cell strand sequencing and long reads | Nature Biotechnology

Scaffolding and completing genome assemblies in real-time with nanopore sequencing | Nature Communications

Scaffolding and correcting physical map contigs based on the genome map... | Download Scientific Diagram

Types of assembled contigs and alignments to REF contigs. a A contig... | Download Scientific Diagram

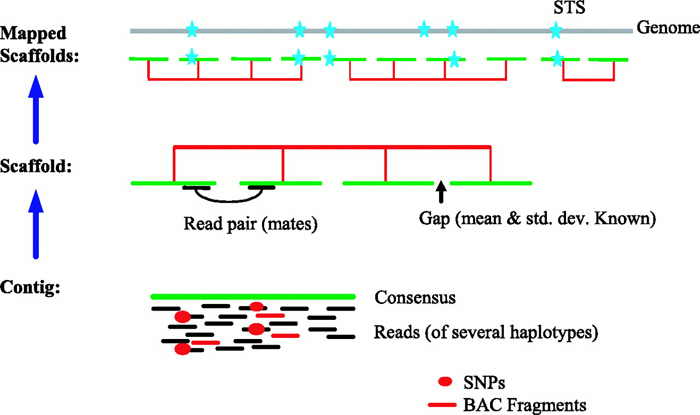
Whole Genome Sequencing Process of Creating Contigs from Fragment Reads | Download Scientific Diagram

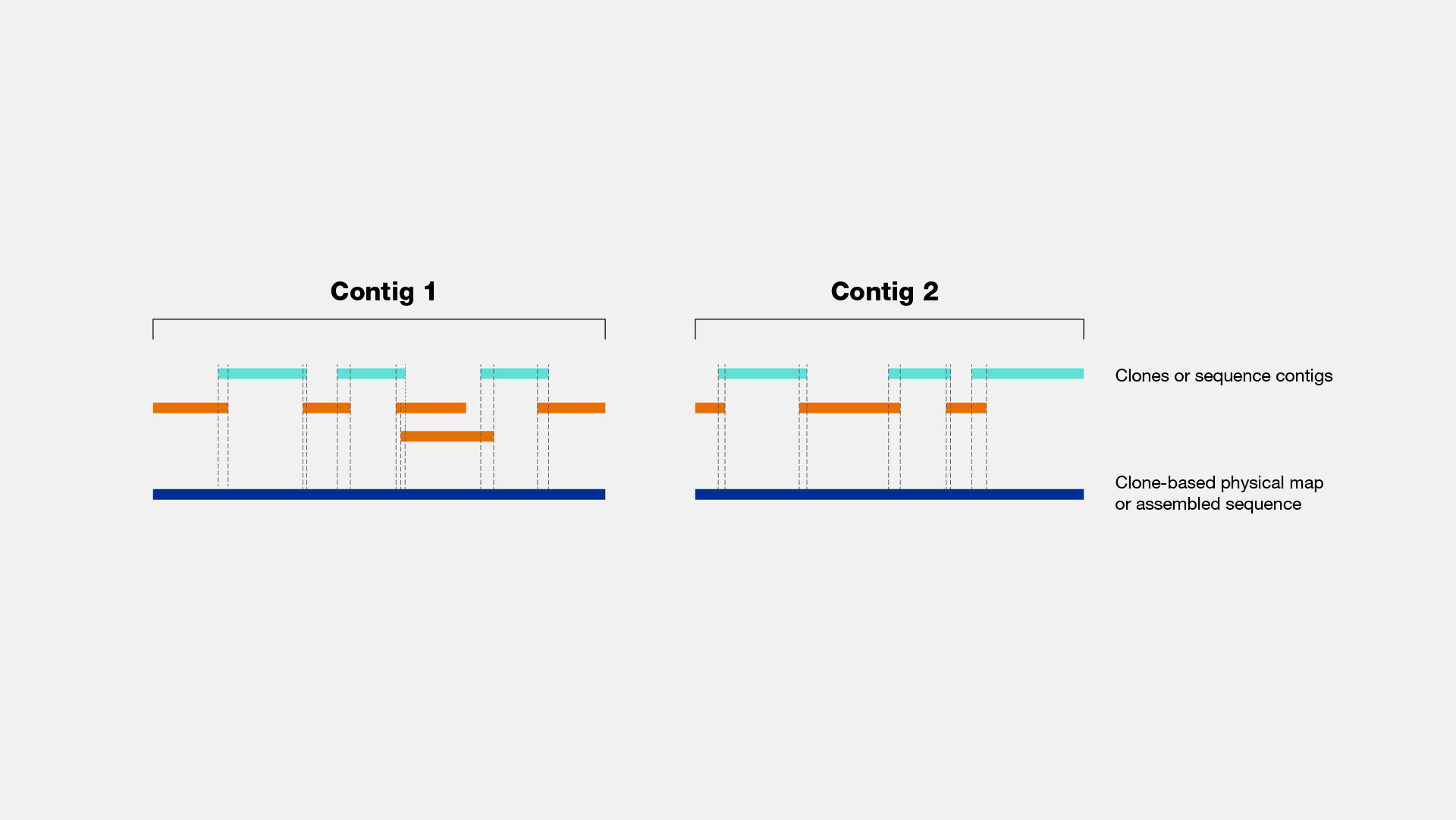
Levels of clone and sequence coverage. A `®ngerprint clone contig' is... | Download Scientific Diagram
![PyDamage: automated ancient damage identification and estimation for contigs in ancient DNA de novo assembly [PeerJ] PyDamage: automated ancient damage identification and estimation for contigs in ancient DNA de novo assembly [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/11845/1/fig-1-full.png)
PyDamage: automated ancient damage identification and estimation for contigs in ancient DNA de novo assembly [PeerJ]